
- KOR
- |
- ENG
Microbiome Service
Welcome to eGnome, where we proudly present our state-of-the-art 3rd generation microbiome analysis technology. To facilitate understanding, we employ a straightforward supermarket analogy.
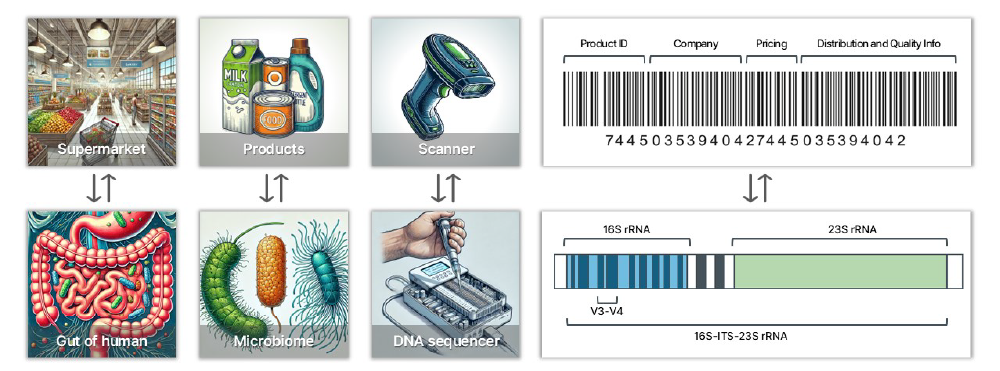
Consider a supermarket where you wish to ascertain the variety and quantity of products available. By scanning the barcodes on each item, you can determine the manufacturing company and the specific product. This method allows you to compile comprehensive data on the types and quantities of products from each company.
Applying this analogy to our microbiome analysis, we examine the gut microbiome (akin to the supermarket’s products) by analyzing the rRNA operon (comparable to barcodes) of each microorganism. The rRNA operon comprises the 16S, ITS, and 23S regions. Reading these regions is analogous to scanning barcodes to gather detailed information. Most microbiome healthcare companies utilize 2nd generation sequencing, which reads a limited portion of the 16S region (approximately 430 base pairs), providing genus-level information. In contrast, eGnome’s 3rd generation method reads the complete 16S-ITS-23S operon, yielding species-level information. This approach provides approximately 4,300 base pairs of data, offering ten times more information than the 2nd generation method.
For instance, take the probiotic Lactobacillus acidophilus. 'Lactobacillus' represents the genus, while 'acidophilus' denotes the species. In our supermarket analogy, the genus corresponds to the company name, and the species corresponds to the product name. A single genus can encompass over 100 different species.

Species-level identification offers 100 times greater resolution, allowing precise differentiation between beneficial and harmful bacteria. While some assert that 2nd generation methods can identify species, their accuracy is merely around 25%, rendering them unreliable. eGnome’s method boasts a 99.9% accuracy in species identification. We employ the 3rd generation sequencer from Oxford Nanopore Technology to achieve precise species-level resolution. After seven years of research, we finalized our EG Mirror technology in 2022 and published our findings in the journal Microbiology Spectrum. By 2024, we launched version 2, now acknowledged as the most accurate commercially available platform.
Leveraging this advanced technology, eGnome offers a diverse array of microbiome services. Our current portfolio includes gut microbiome analysis (EG gut PRO), vaginal microbiome analysis (EG vaginal PRO), and pet gut microbiome analysis (EG pet Dog, Cat). Future services will encompass skin microbiome analysis, children’s growth and development microbiome analysis, obesity microbiome analysis, and senior microbiome analysis.

